Pistol
Rfam ID: RF02679
click into different sections:
Timeline
-
2015 Discovery, Secondary structure[1]
-
2015 Biochemical analysis of pistol ribozyme[2]
-
2016 The pseudoknot fold facilitating sitespecific in-line cleavage[3]
-
2017 Crystal structure of Pistol shows an evolutionarily conserved cleavage mechanism that is like other self-cleaving ribozymes[4]
-
2017 Structural and Catalytic Roles for an Active-Site Adenosine and Hydrated Mg(2+) in Pistol Ribozymes[6]
-
2019 Comparison of the Structures and Mechanisms of the Pistol and Hammerhead Ribozymes[7]
-
2020 Crucial Roles of Two Hydrated Mg2+ Ions in Reaction Catalysis[8]
-
2020 Mg2+ -dependent cleavage of a pistol ribozyme reveals a fail-safe secondary ion for catalysis[9]
-
2021 Biochemical analysis of cleavage and ligation activities of the pistol ribozyme from Paenibacillus polymyxa[12]
-
2022 New classical and combined quantum mechanical/molecular mechanical simulation of pistol ribozyme[13]
Description
The pistol ribozyme is an RNA structure that catalyzes its own cleavage at a specific site. In other words, it is a self-cleaving ribozyme. The pistol ribozyme was discovered through comparative genomic analysis. Subsequent biochemical analysis determined further biochemical characteristics of the ribozyme. This understanding was further advanced by an atomic-resolution crystal structure of a pistol ribozyme.
Structure and mechanism
2D representation
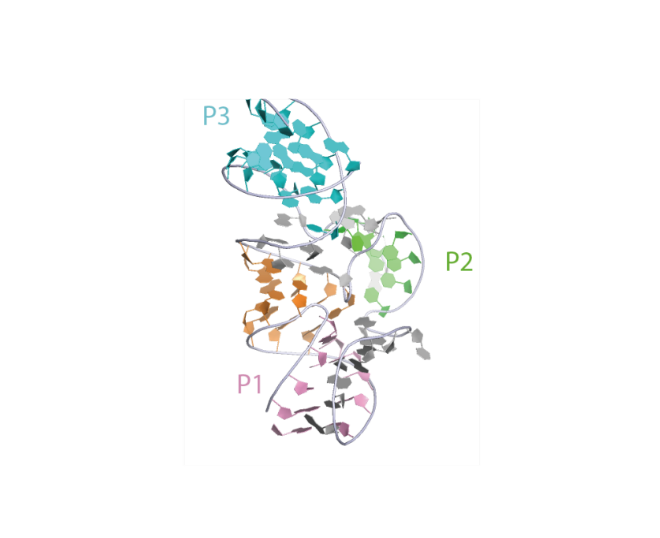
Secondary structure of the Pistol ribozyme. The sequence is color coded according to helical segments observed in the tertiary structure.

|
3D visualisation
Crystal structure of the Pistol ribozyme. This representation was generated from PDB ID: 6R47 at 3.1 Å resolution. Helices are differentiated by color.
 |
|
Catalytic centre
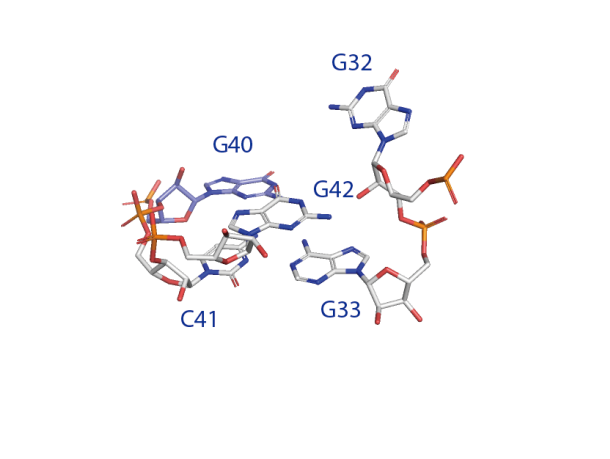
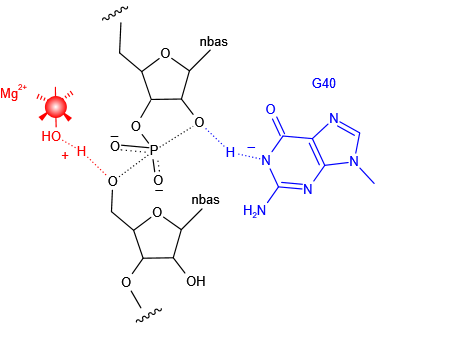
(left) Close view of the catalytic center of the ribozyme. G40 N1 is hydrogen bonded to the O2' nucleophile and acts as the general base in the cleavage reaction. (right) Proposed mechanisms for pistol ribozyme.Metal ions as the general base.
 |
 |
Chemical mechanism
The secondary and 3D structures of the pistol and hammerhead ribozymes have a number of similarities, suggesting that they might have related catalytic mechanisms [7]. Structural comparison reveals that the environment of G40 in pistol is very similar to that of G12 in the hammerhead, with G40 N1 2.5Å from the O2' nucleophile, so that G40 is the likely general base for the cleavage reaction. The rate of cleavage rises log-linearly with pH up to pH = 8.75 and then reduces at higher pH values. Substitution of A G40I modification shifted down the pH maximum by ~0.5 pH units [7], implicating G40 in proton transfer in the transition state. If pistol and the hammerhead ribozymes were mechanistically identical we would expect that the general acid would be a 2’-hydroxyl group, but this is not supported by experimental data. First, in the crystal structure the nearest O2' (A32) is 5 Å from the O5' [7]. Second, an A32 O2' NH2 substitution led to a 100-fold reduction of cleavage rate, but the pH profile was essentially unaltered from the unmodified ribozyme, in marked contrast to the equivalent data from the hammerhead ribozyme [7]. So rather than using the A32 O2' group it seems most probable that the pistol ribozyme uses an inner sphere water molecule of a bound Mg2+ ion as general acid. The rate of cleavage was found to increase as the divalent metal ion was substituted Ca2+ < Mg2+ < Mn2+ < Co2+, and there was a log-linear dependence on the pKa of the hydrated metal ion [7]. This is consistent with the metal ion acting as the general acid in the cleavage reaction, so that the rate increases with the reactivity of the ion as its pKa is lowered. At present this is the only ribozyme that appears to use a hydrated metal ion as a general acid in the cleavage reaction.
References
[1] New classes of self-cleaving ribozymes revealed by comparative genomics analysis.
Weinberg, Z., Kim, P. B., Chen, T. H., Li, S., Harris, K. A., Lunse, C. E. & Breaker, R. R.
Nature Chemical Biology, 11, 606-10.(2015)
[2] Biochemical analysis of pistol self-cleaving ribozymes.
Harris, K. A., C. E. Lunse, S. Li, K. I. Brewer and R. R. Breaker
RNA 21(11): 1852-8.(2015)
[3] Pistol ribozyme adopts a pseudoknot fold facilitating site-specific in-line cleavage.
Ren, A., Vusurovic, N., Gebetsberger, J., Gao, P., Juen, M., Kreutz, C., Micura, R. & Patel, D. J.
Nature Chemical Biology, 12, 702-8.(2016)
[4] Crystal structure of Pistol, a class of self-cleaving ribozyme.
Nguyen, L. A., Wang, J. & Steitz, T. A.
Proc Natl Acad Sci U S A, 114, 1021-6.(2017)
[5] Deep Sequencing Analysis of Aptazyme Variants Based on a Pistol Ribozyme.
Kobori, S., K. Takahashi and Y. Yokobayashi
ACS Synth Biol 6(7): 1283-1288.(2017)
[6] Atom-Specific Mutagenesis Reveals Structural and Catalytic Roles for an Active-Site Adenosine and Hydrated Mg(2+) in Pistol Ribozymes.
Neuner, S., C. Falschlunger, E. Fuchs, M. Himmelstoss, A. Ren, D. J. Patel and R. Micura
Angew Chem Int Ed Engl 56(50): 15954-15958.(2017)
[7] Comparison of the Structures and Mechanisms of the Pistol and Hammerhead Ribozymes.
Wilson, T. J., Y. Liu, N. S. Li, Q. Dai, J. A. Piccirilli and D. Lilley
J Am Chem Soc 141(19): 7865-7875.(2019)
[8] Crucial Roles of Two Hydrated Mg(2+) Ions in Reaction Catalysis of the Pistol Ribozyme.
Teplova, M., Falschlunger, C., Krasheninina, O., Egger, M., Ren, A., Patel, D. J. & Micura, R.
Angew Chem Int Ed Engl, 59, 2837-43.(2020)
[9] Molecular dynamics analysis of Mg(2+) -dependent cleavage of a pistol ribozyme reveals a fail-safe secondary ion for catalysis.
Joseph, N. N., R. N. Roy and T. A. Steitz
J Comput Chem 41(14): 1345-1352.(2020)
[10] Fundamental studies of functional nucleic acids: aptamers, riboswitches, ribozymes and DNAzymes.
Micura, R. and C. Hobartner
Chem Soc Rev 49(20): 7331-7353.(2020)
[11] Circularly-Permuted Pistol Ribozyme: A Synthetic Ribozyme Scaffold for Mammalian Riboswitches.
Mustafina, K., Y. Nomura, R. Rotrattanadumrong and Y. Yokobayashi
ACS Synth Biol 10(8): 2040-2048.(2021)
[12] Biochemical analysis of cleavage and ligation activities of the pistol ribozyme from Paenibacillus polymyxa.
Lihanova, Y. and C. E. Weinberg
RNA Biol 18(11): 1858-1866.(2021)
[13] Who stole the proton? Suspect general base guanine found with a smoking gun in the pistol ribozyme.
Ekesan, S. and D. M. York
Org Biomol Chem.(2022)
 Home
Home Database
Database Research
Research About us
About us

